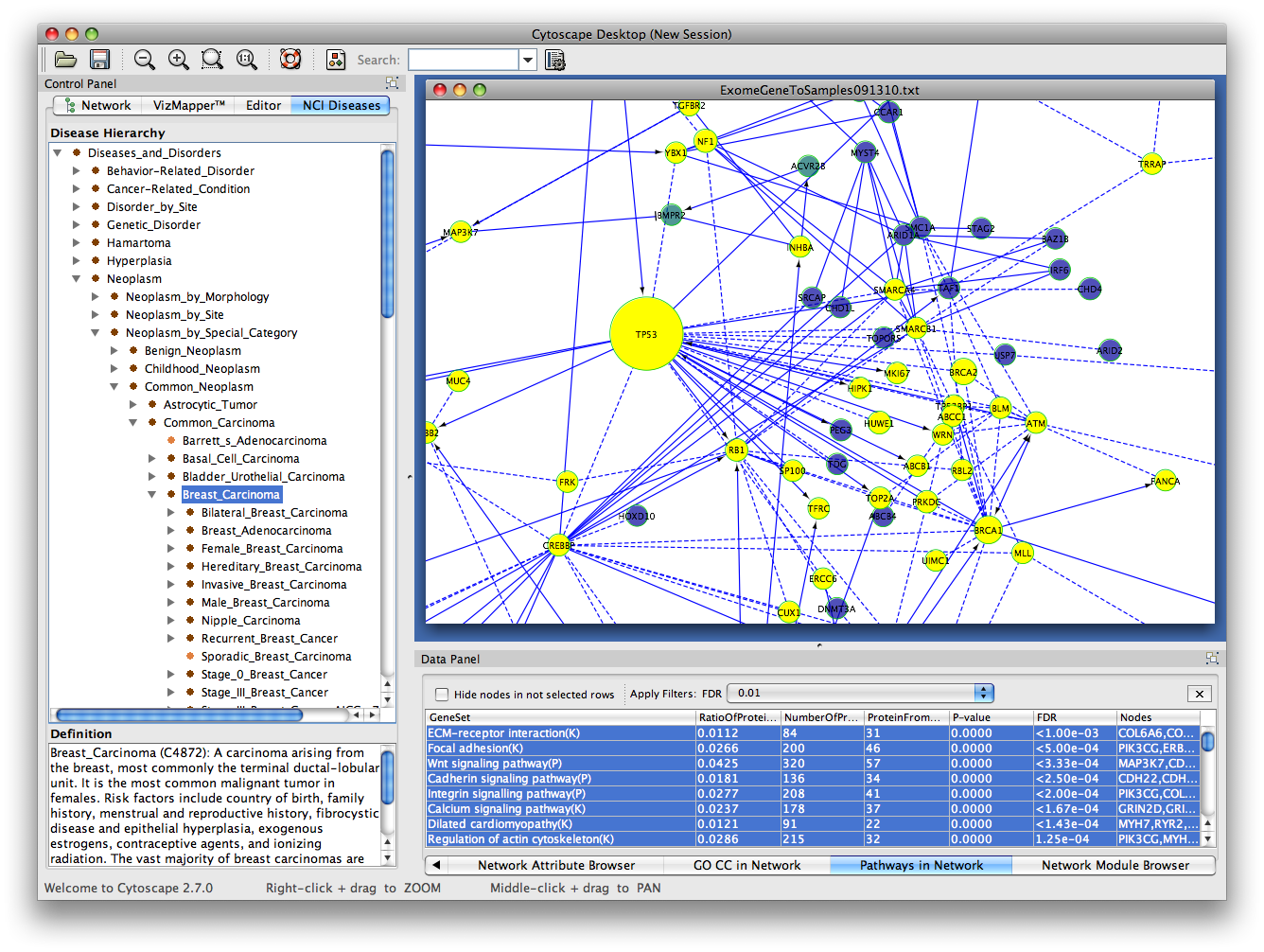
There are two different types of clustering algorithms supported by clusterMaker: Will be available in a session that was saved after clustering. Hierarchical or k-Means clusters if either of those methods had beenīecause information about clusters is saved inĬytoscape attributes, the Eisen TreeView and Eisen KnnView options (without clustering), and options appropriate for displaying This will bring up the settings dialog for the selected algorithmĪ subset of the visualization options, including showing a heat map of the data Where algorithm is the clustering algorithm you wish to use (see Figure 2). New Cluster menu hierarchy under the Plugins main menu.Įach of the supported clustering algorithms appears as a separate Once clusterMaker is installed, it will install a Select clusterMaker and click the Install button.įigure 2. Select Analysis under Available for Install. Is available in the Analysis group of plugins. You must be running Cytoscape 2.8.2 or newer. To download clusterMaker using the plugin manager, Create New Network with Nested Networks from AttributeĬlusterMaker is available through the Cytoscape plugin manager orīy downloading the source directly from the Cytoscape svn repositoryĬytoscape Subversion Server information, or browse the csplugins/ucsf/scooter/clusterMaker.SCPS (Spectral Clustering of Protein Sequences).Gene expression analysis in a network context.įinding complexes in proteomic and genetic interaction data.įunctional annotation by clustering protein similarity networks. In the BMC Bioinformatics publication: " clusterMaker: A Multi-algorithm Clustering Plugin for Cytoscape", currently submitted. The other three tutorials describe the steps necessary to reproduce the scenarios described The first tutorial:Ĭluster Maker covers some of the basic featuresĪnd uses of clusterMaker. Open Tutorials web site under the "Cytoscape Tutorials" section. In addition to this documentation, there are four tutorials available on the "meta nodes" to allow interactive exploration of the putative familyĪssociations within the Cytoscape network, and results may alsoīe shown as a separate network containing only the intra-cluster edges, or with inter-cluster edges added back.ĬlusterMaker requires version 2.8.2 or newer of CytoscapeĪnd is available from the Cytoscape plugin manager under Hierarchical, k-medoid, AutoSOME, and k-means clusters may beĭisplayed as hierarchical groups of nodes or as heat maps.Īll of the network partitioning cluster algorithms create collapsible MCODE, community clustering (GLAY), SCPS, and AutoSOME for partitioning networks based on similarity K-means for clustering expression or genetic data and MCL, transitivity clustering, affinity propagation, Current clustering algorithms include hierarchical, k-medoid, AutoSOME, and UCSF clusterMaker is a Cytoscape plugin that unifiesĭifferent clustering techniques and displays into a single In this screenshot, theĮxpression data in the sampleData file galFiltered.cys has been clustered using the hierarchical methodĪnd displayed as a heatmap with associated dendrogram. Finally, to enrich the visual exploration, it is possible to visually render local topological properties of the network (e.g. The plugin also allows the extraction of parts of the network that contain a selected subset of reactions. the organism or perturbation under study), we propose facilities to edit/select putative biochemical transformations. Since the definition of this list is closely related to experimentation (i.e. Inference requires a list of potential biochemical transformations. Here we present a new plugin for Cytoscape dedicated to the inference and visualization of high-resolution metabolomic networks. There is currently no available software that allows inference and visualization of such high-resolution metabolomic networks directly from raw data. To analyze, explore and interpret these two kinds of relations, powerful visualization tools are required. The combination of these two inference methods generates networks containing hundreds of nodes (metabolites) and hundreds of predicted edges (biochemical reactions and/or high correlations). 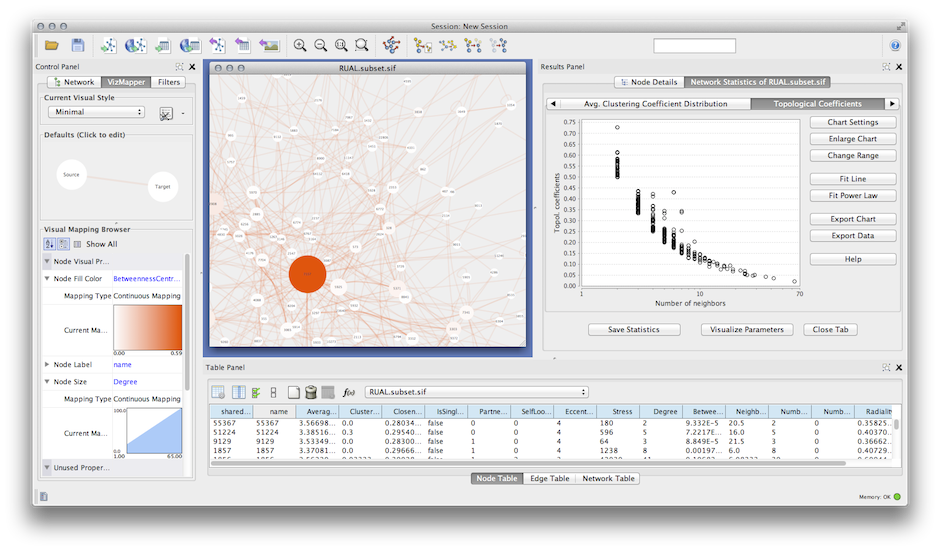
Moreover, perturbation studies allow the use of correlation analysis to infer/confirm links between metabolites that correlate across various conditions. Such high-resolution data has also been used to predict ab initio biochemical interactions between metabolites. Recently, ultra high-resolution mass spectrometry (FTICR-MS or Orbitrap) has been successfully used in metabolomic studies. Various spectrometric technologies are capable of identifying thousands of metabolites. Metabolomics aims at the identification and quantification of all metabolites that are present in a cell, tissue or biofluid at a given moment and under particular conditions.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
